scOrange Blog
-
Categories:
- clustering
- t-sne
- marker-genes
- tsne
- annotator
- data-mining
- embedding
- embryo
- genomics
- markers
Author: Martin Stražar, Nov 23, 2018
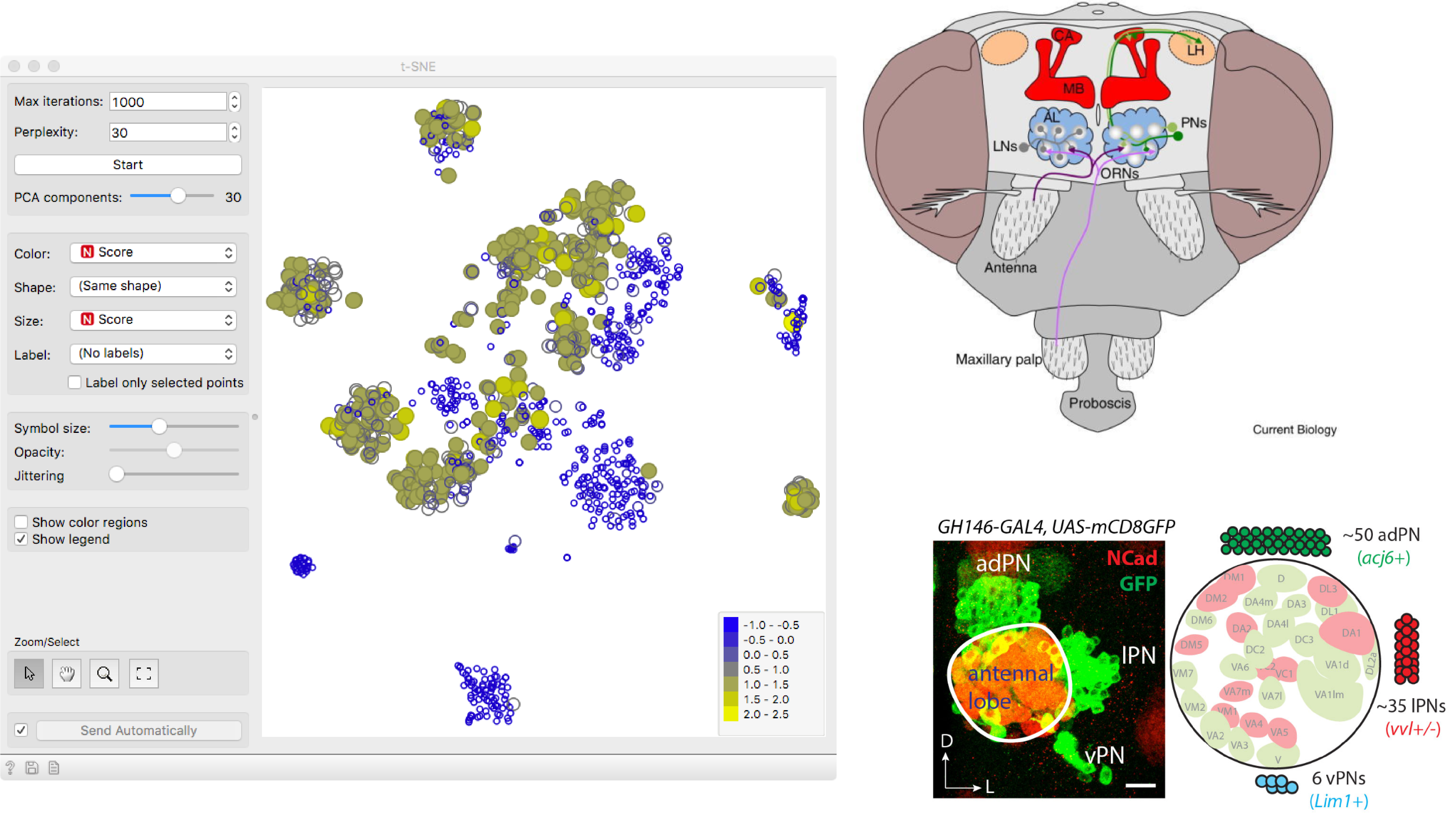
Exploring the Fruit Fly Olfactory System
Explore different subsets of cells and related them to known biological processes.
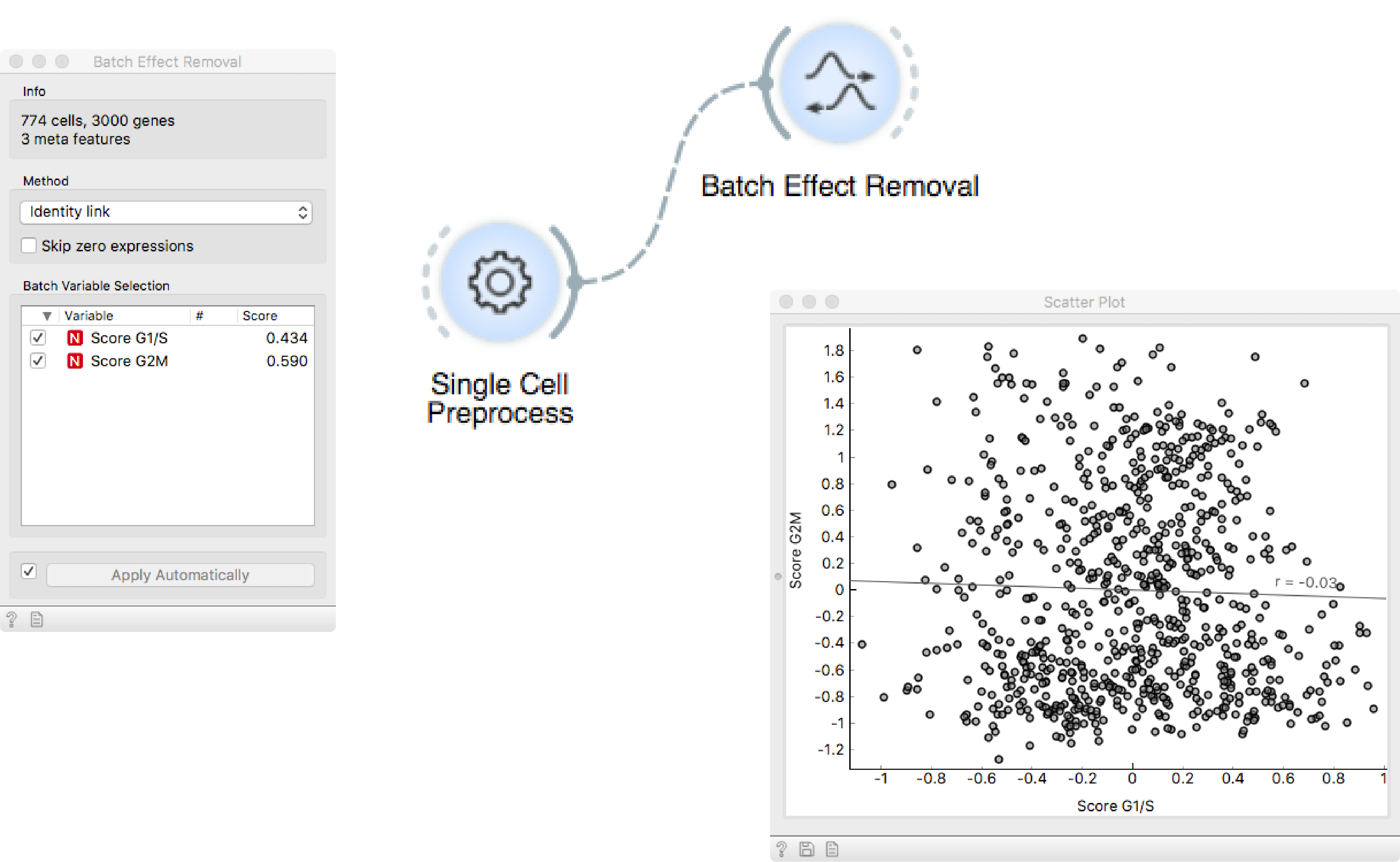
Author: Martin Stražar, Oct 5, 2018
Nuts and bolts: Preprocessing and Removing Batch Effects
Preprocess raw read count data to make expression values comparable across cells and genes. Use Gene Markers and Score Cells to estimate the cell cycle phase of each cell and regress out this confouding factor.